BLAST-XYplot viewer
A Massive BLAST Visualization and Analysis Tool for
whole-genome sequenced bacteria
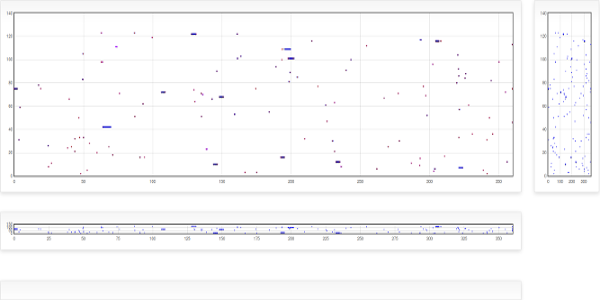
Welcome to BLAST-XYplot viewer, a tool programed in Perl which performs a massive and systematic BLAST search of a desired group of proteins or genes against selected bacterial genomes. In order to handle high number of BLAST-results, these are depicted in an x,y plot where the x-value corresponds to a "relative position" calculated for each gene into the genome and the y-value corresponds to the number of bacterial replicon (chromosome or plasmid). In this representation scheme, the circular replicon is projected as a line with fixed length of 360 which are the degrees in a circle, giving a delimited space where any replicon can be represented independently of their actual size and each gene can be mapped by their Relative Position (0-360) instead of their real (nucleotide) position. The plot allows a rapid and easy visualization of the closely positioned genes between thousands of results. In addition, a table of data containing the information of the subjects found is scored, sorted and written to a file that could be analyzed in a spreadsheet. In a typical run, this tool can perform the search of twenty sequences on one hundred of replicons in several minutes and search of one hundred sequences on all bacterial genomes takes a few hours.
If you use results obtained with this tool, please cite as:
Pedraza-Pérez, Y., Cuevas-Vede, R.A., Canto-Gómez, Á.B., López-Pliego, L., Gutiérrez-Ríos, R.M., Hernández-Lucas, I., Rubín-Linares, G., Martínez-Laguna, Y., López-Olguín, J.F. and Fuentes-Ramírez, L.E., 2018. BLAST-XYPlot Viewer: A Tool for Performing BLAST in Whole-Genome Sequenced Bacteria/Archaea and Visualize Whole Results Simultaneously. G3: Genes, Genomes, Genetics, 8: 2167-2172; https://doi.org/10.1534/g3.118.200220

